Introduction
Traditional metabolomics can reflect changes in metabolites within an organism and identify potentially activated pathways. However, since a single metabolite may participate in multiple metabolic pathways without a change in its total abundance, this approach has limitations. Metabolic networks are complex and dynamic, whereas standard metabolomics provides only static information on metabolite abundance. Isotope tracing effectively addresses these limitations.
Stable isotope-labeled analysis (MFA) is particularly capable of capturing the dynamic changes within cellular metabolic networks, reflecting metabolic reprogramming patterns under genetic or environmental perturbations. By adding specifically labeled substrates (e.g., 13C or 15N) to the culture medium, these isotopes are incorporated into the biological system. As cells grow, they absorb and metabolize these substrates into various downstream metabolites. Detecting labeled metabolites in the samples allows researchers to trace and characterize cellular metabolic processes.
BIOTREE has established an untarget isotope tracing method based on the LC-MS/MS platform. Unlike target isotope tracing, which replies on metabolite standards to identify labeled metabolites and limits the number of labeled metabolties detection, untarget isotope tracing detects labaled metabolites from identifications of untarget metabolomics screening, and detects hundreds of labeled metabolites.
Wang, R., Yin, Y., Li, J. et al. Global stable-isotope tracing metabolomics reveals system-wide metabolic alternations in aging Drosophila. Nat Commun 13, 3518 (2022).
Technical Advantages
◉ Accurate: Aligned with international MSI (Metabolomics Standards Initiative) standards, achieving a Level 1 identification accuracy of up to 98.5%.
◉ Rigorous: Implements a standardized database construction workflow and an ultra-strict 6D quality control system.
◉ Comprehensive: Features the BiotreeDB 4.0 with over 20,000 standards, complemented by exclusive proprietary databases for clinical, plant, and lipid research.
◉ High Yield: Delivers high detection rates, with an average of 800+ Level 1 identified metabolites per sample.
◉ Extensive: Offers broad tracking coverage, capable of simultaneously tracing stable isotope labels across thousands of metabolites.
◉ Superior: Utilizes high-sensitivity, high-resolution instrumentation with exceptional performance stability.
Sample Requirement
Liquid | 100ul/sample |
Tissue | 100mg/sample |
Organoid | 100mg/sample |
Cells | 1*10^7 cells/sample |
LC-MS Platform
Orbitrap Exploris 120, Thermo
Applications
◉ Broad Applications: Untargeted metabolic flux analysis is widely applicable in life sciences and pharmaceutical research, spanning fields such as cellular metabolic regulation, discovery of novel metabolic pathways, disease metabolic mechanisms, drug target discovery and validation, drug efficacy and toxicity evaluation, diagnostic and prognostic biomarkers, pharmacometabolomics, and precision medicine.
◉ Genetic Engineering: Optimizing target metabolite yields in engineered strains and characterizing metabolic functional changes before and after genetic modification.
◉ Disease Mechanisms: Elucidating the mechanisms behind personalized therapy with tumor metabolic inhibitors and identifying biomarkers for the early diagnosis of disease onset and progression.
◉ Metabolic Reprogramming: Investigating mechanisms of immune metabolic reprogramming in inflammatory macrophages and chloroplast metabolic reprogramming in plants.
Example Publication
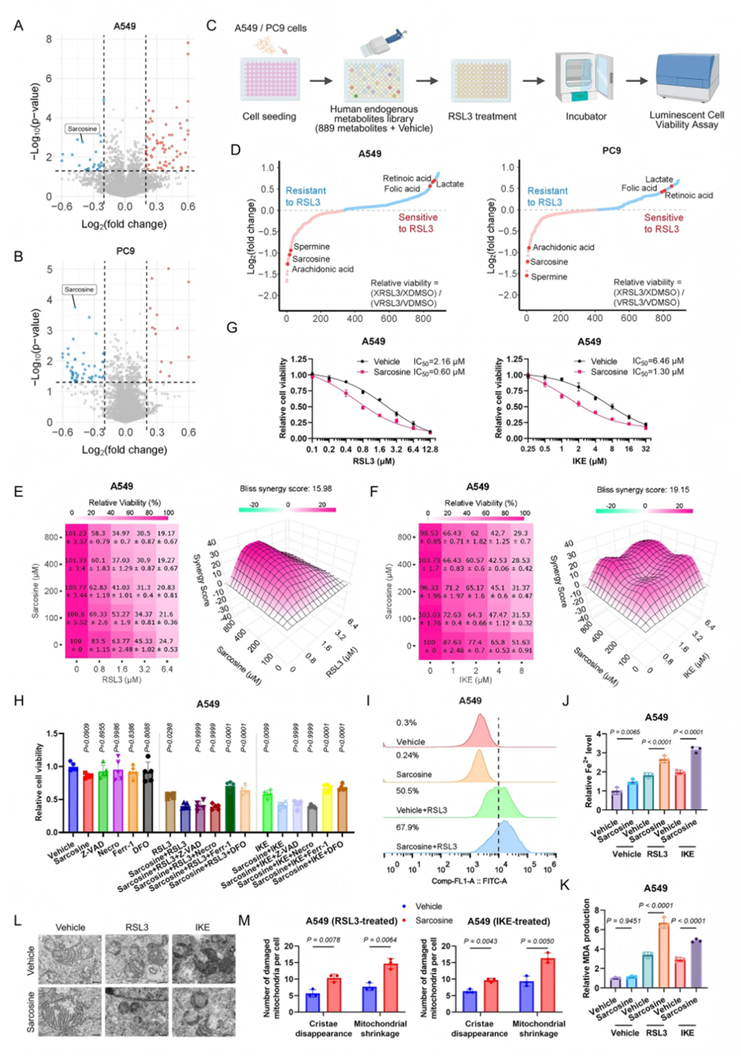
Metabolite Library Screening Reveals the Association Between Sarcosine and Ferroptosis
Title: Sarcosine Sensitizes Lung Adenocarcinoma to Chemotherapy by Dual Activation of Ferroptosis via PDK4/PDHA1 Signaling and NMDAR-Mediated Iron Export
Journal: Experimental Hematology & OncologyImpact Factor: 13.5
Research Background:
Lung cancer remains the leading cause of cancer-related deaths globally, with Lung Adenocarcinoma (LUAD) being the most common subtype of non-small cell lung cancer (NSCLC). Despite advancements in treatment, the five-year survival rate remains below 26%, largely due to chemotherapy resistance. Ferroptosis, an iron-dependent form of regulated cell death driven by lipid peroxidation, has emerged as a promising strategy to overcome therapeutic resistance. While evidence suggests that modulating ferroptosis can enhance chemosensitivity, the precise mechanisms for inducing ferroptosis and its metabolic regulation remain unclear. Sarcosine, an endogenous metabolite linked to invasion and metastasis in cancers like prostate cancer, has an unknown role in LUAD.
This study utilized metabolomics screening to discover that sarcosine significantly enhances the sensitivity of LUAD cells to ferroptosis inducers (FINs). Integrating clinical data with mechanistic studies, the research confirms that sarcosine regulates ferroptosis through a dual mechanism. Furthermore, the combination of sarcosine and cisplatin synergistically improves chemosensitivity. Clinically, high sarcosine levels in patient samples correlate with improved prognosis. This study fills a critical gap regarding the role of sarcosine in ferroptosis and LUAD, offering a new target to overcome chemotherapy resistance. Given its safety profile and metabolic regulatory properties, sarcosine holds significant potential as an adjuvant for platinum-based chemotherapy.
Key Findings:
Metabolite Library Screening Reveals Sarcosine Enhances Ferroptosis Sensitivity in LUAD Cells
To identify key metabolites regulating ferroptosis and verify whether sarcosine enhances LUAD cell sensitivity through specific mechanisms, the study first performed untargeted metabolomics on LUAD cell lines (A549 and PC9) treated with the ferroptosis inducer RSL3. This analysis revealed unique metabolic signatures, including a significant downregulation of sarcosine (Fig. 1A-B). Subsequently, screening a library of 889 human endogenous metabolites in an RSL3-induced ferroptosis model identified sarcosine as a potent enhancer of RSL3-mediated ferroptosis (Fig. 1C-D).
◉ Specificity of Action: Cytotoxicity assays demonstrated that combining sarcosine with ferroptosis inducers (e.g., RSL3, IKE) significantly reduced cell viability. This synergistic effect was completely abolished by ferroptosis inhibitors—specifically the lipid peroxidation scavenger Ferrostatin-1 (Fer-1) and the iron chelator Deferoxamine (DFO). Conversely, inhibitors of apoptosis (Z-VAD-FMK) or necroptosis (Necrosulfonamide) had no effect, confirming the specificity of sarcosine’s pro-ferroptotic activity (Fig. 1E–H).
◉ Mechanistic Markers: Further analysis showed that exogenous sarcosine significantly amplified lipid reactive oxygen species (Lipid-ROS) generation (Fig. 1I), intracellular ferrous iron ( Fe2+Fe2+ ) accumulation (Fig. 1J), and malondialdehyde (MDA) levels (Fig. 1K) induced by RSL3.
◉ Ultrastructural Changes: Transmission electron microscopy (TEM) revealed characteristic ferroptotic ultrastructural alterations following sarcosine treatment, including the disappearance of mitochondrial cristae and mitochondrial shrinkage (Fig. 1L).
Conclusion: Sarcosine significantly enhances the sensitivity of LUAD cells to ferroptosis by promoting iron-dependent lipid peroxidation and inducing mitochondrial dysfunction.
