Introduction
Orbitrap Astral Zoom is the up-to-date MS launched by Thermo Fisher, and provides unprecedented improvements in scan speed, sensitivity, and dynamic range.
Data-Independent Acquisition (DIA) mode is used for untarget proteomics testing. Within a predefined scanning window, this approach fragments all precursor ions, offering superior reproducibility and generating richer spectral information. Consequently, this significantly enhances both the coverage and reliability of qualitative and quantitative results.
Incorporating magnetic bead-based sample pretreatment, Biotree's DIA proteomics provides unbiased adsorption of all proteins. This solution not only ensures the stability and consistency of identification results but also markedly reduces the required volume of biological samples. Ultimately, it provides efficient and robust technical support for advanced proteomic analysis.
Technical Advantages
◉ Fast Scanning Speed Throughout the LC-MS/MS workflow, the system acquires 100–150 high-resolution MS/MS spectra per second while maintaining a resolution of 40,000–50,000. This exceptional speed makes it particularly ideal for large-scale clinical cohort studies, significantly shortening analysis cycles.
◉ High Accuracy By combining high-precision mass spectrometry acquisition modes with proprietary algorithmic corrections, the platform achieves superior accuracy in both protein identification and quantification, substantially minimizing detection bias.
◉ High Sensitivity The enhanced scanning speed translates directly into improved sensitivity, enabling the identification of low-abundance proteins from minimal sample inputs.
◉ Exceptional Stability Designed for robustness, the system maintains consistent performance even during the continuous analysis of thousands of samples, ensuring reliable data quality across extensive runs.
Sample Requirement
Tissue | 10mg/sample |
Cell/Bacteria | 3*10^6/sample |
Fresh plant tissue | 400mg/sample |
Plant fruit/seed | 4g/sample |
LC-MS Platform
Orbitrap Astral Zoom,Thermo
Applications
◉ Food Science: Research aimed at enhancing food safety, improving texture, and optimizing nutritional value.
◉ Animal Husbandry: Studies focused on improving meat quality and optimizing breeding conditions.
◉ Virology: Discovery and characterization of novel viral inhibitory factors.
◉ Agriculture: Investigation into the molecular mechanisms governing crop development.
◉ Disease Mechanisms: Elucidation of the molecular pathways underlying biological diseases.
◉ Biomarker Discovery: Identification and validation of biomarkers for various diseases.
◉ Drug Target Identification: Research dedicated to discovering and validating therapeutic drug targets.
◉ Drug Mechanism of Action: Studies exploring the specific mechanisms by which drugs exert their effects.
Example Publication
Clinical:
4D-Based Serum Proteomics Analysis
Title: High-resolution serum proteome trajectories in COVID-19 reveal patient-specific seroconversion
Journal: EMBO Molecular Medicine
Impact Factor: 14.26
Methodology: Protein Corona-DIA Quantitative Proteomics
Study Overview:
The discovery of valuable biomarkers using proteomics is a major focus in current clinical research. This study employed Protein Corona-DIA quantitative proteomics to analyze serum protein profiles in COVID-19 patients compared to PCR-negative controls.
The results revealed that 26% of the detected proteins underwent significant changes during the disease course. Notably, proteins associated with the innate immune system, coagulation regulation, and lipid homeostasis showed substantial alterations. Furthermore, the study identified ITIH4 as a novel potential biomarker. These findings provide critical insights into the host protein regulatory mechanisms underlying SARS-CoV-2 infection and disease progression.
Agricultural:
Dietary Flaxseed Oil Modulates Hepatic Protein Composition
Title: Coral and its symbionts responses to the typical global marine pollutant BaP by 4D-Proteomics approach
Journal: ENVIRONMENTAL POLLUTION
Impact Factor: 9.988
Methodology: Protein Corona-DIA Quantitative Proteomics (combined with 16S rRNA sequencing)
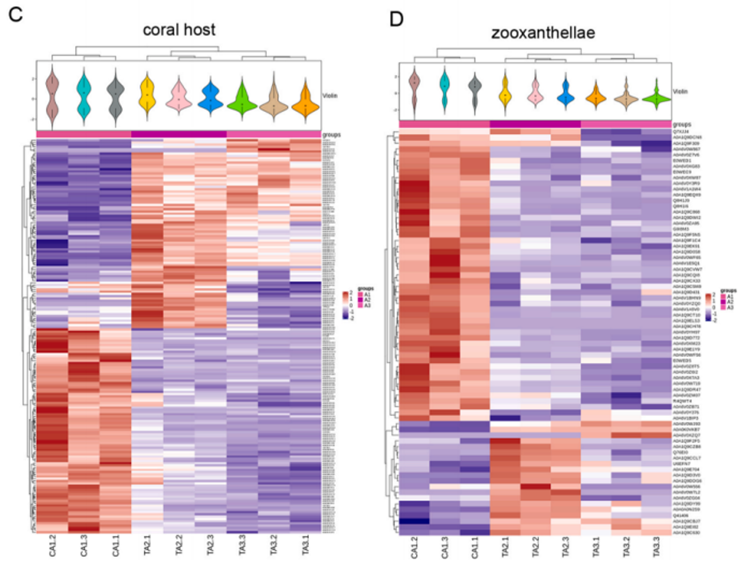
Study Overview: The mutualistic symbiosis between reef-building corals and Symbiodiniaceae (zooxanthellae) is the cornerstone of coral reef ecosystems, yet this relationship is increasingly threatened by marine pollution. Benzo[a]pyrene (BaP), a globally distributed marine environmental pollutant, poses a significant risk to marine ecosystems. However, the stress responses of coral holobionts to such global pollutants remain poorly understood.
This study focused on the stony coral Acropora formosa, utilizing Protein Corona-DIA quantitative proteomics alongside 16S rRNA sequencing to investigate its response to BaP stress. The results elucidate the toxicological mechanisms of coral responses to BaP exposure at the protein level. These findings provide a crucial scientific basis for the conservation and restoration of coral reef ecosystems.
